In vitro retinal disease modelling service
Retinal disease modelling development service: this service enables the development of isogenic human induced pluripotent stem cells (iPSCs) derived retinal organoids and Retinal Pigment Epithelium (RPE) from patient or gene edited lines. This service from Newcells Biotech is designed to support retinal and eye research and is offered as part of our retina models service.
Targeted mutations can model retinal diseases, more specifically monogenic inherited retinopathies such as retinitis pigmentosa (RP), Stargardt disease, Usher’s syndrome and Leber congenital amaurosis.
These retinal models are fully characterised and can then be used to carry out efficacy and safety testing of novel compounds or gene therapy vector assessments for retinal therapies. The follow-on studies using retinal disease modelling enable the comparison between WT and mutant phenotypes after differentiation into retinal organoids or Retinal Pigment Epithelium (RPE) giving you insights into the basis of the retinal disease or therapeutic mechanism.
Service outputs
- Morphological characterisation (Brightfield imaging)
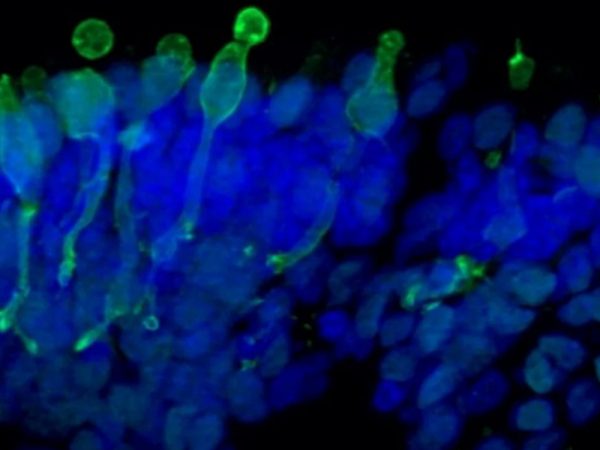
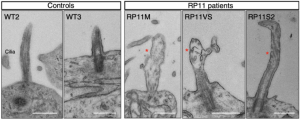
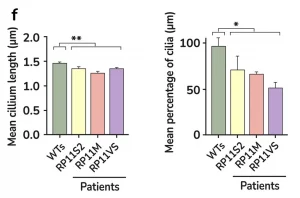
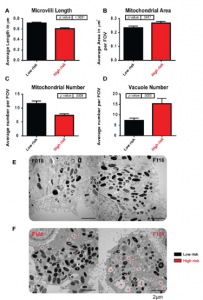
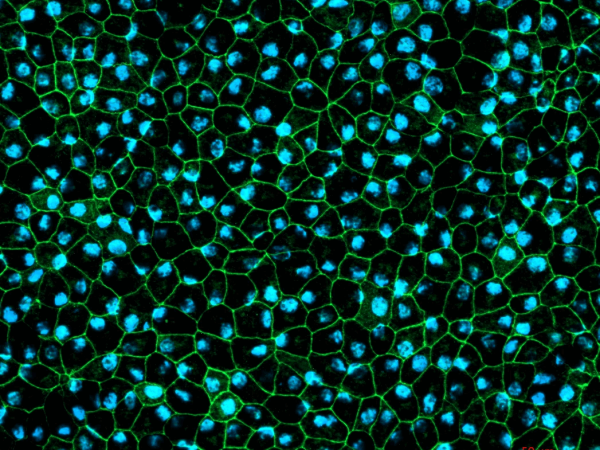
- Key markers analysis (Immunofluorescence, TEM or SEM and RT-PCR)
- Phagocytosis of photoreceptor outer segment (RPE only)
- TEER (RPE only)
- Growth factor secretion (RPE only)
- Protein expression (RPE only)
- Comparison with heathly donor control